Introduction
As the most common hereditary kidney disease, Autosomal Dominant Polycystic Kidney Disease (ADPKD) is characterized by progressive dilation of bilateral renal cysts [1], leading to the destruction of normal renal parenchyma and progressive decline in renal function, ultimately resulting in End-Stage Renal Disease (ESRD) [2]. Currently, clinical management strategies mainly focus on complication control, and there is still a lack of targeted therapies that can effectively delay disease progression [3] Tovaptan, as the first approved vasopressin V2 receptor antagonist to delay the progression of ADPKD, has been clinically proven to have benefits [4], mainly inhibiting cyst growth mediated by Cyclic Adenosine Monophosphate (cAMP) [5]. Studies have shown that the pathological process of ADPKD involves abnormal activation of multiple signaling pathways, among which the PI3K/Akt and MAPK signaling pathways have been shown to play a central role in regulating abnormal proliferation of cyst epithelial cells [6,7] However, tolvaptan exhibits a wide range of effects beyond single V2 receptor blockade in vivo [8], indicating the existence of a multi-target synergistic mechanism, which is currently not fully understood [9]. This study is based on network pharmacology methods to systematically analyze the mechanism of tolvaptan in delaying the pathological progression of ADPKD. A PPI protein interaction network and a drug target disease network were established, and GO functional annotation and KEGG pathway enrichment analysis were performed. Molecular docking validation was performed on key targets to clarify new targets for ADPKD treatment and provide theoretical basis for clinical drug development.
Materials and methods
Experimental data
The molecular formula, two-dimensional structural formula, and mol format file of tolvaptan were retrieved and obtained through the ChemicalBook chemical database(https://www. ChemicalBook.com/).
Experimental methods
Drug target prediction and standardization: Based on the SwissTarget Prediction platform (http//www. swisstargetprediction.ch/), we predicted the target of action of tolvaptan, combined with the Uniprot database (https://www. uniprot.org/), obtained its potential target of action screen targets with confidence>0 for standardization, and ultimately determined the main target of tolvaptan.
Screening of disease-related genes: Through GeneCards (https://www.genecards.org/), we retrieved ADPKD related target genes from the CTD database using the keyword “Autosomal Dominant Polycystic Kidney Disease” (data updated to April 2024). In the GeneCards database, there is a positive correlation between scoring and disease association. The maximum value of the target gene score obtained is 886.90, the minimum value is 0.73, and the median is 15.95. We selected targets with a score ≥median (15.95) as the main disease targets and merged them with the CTD database results.
Screening of common targets for drug diseases: We used Venny2.1.0 website (https//bioinfogpcnb. csic. es/tools/venny/ Index. html) to conduct intersection analysis of the main targets of tolvaptan and ADPKD, and obtained the common target.
Establishment of drug target disease network: Tovaptan and Tovaptan ADPKD common targets were constructed using Cytoscape 3.10.3 software to construct “Drug target disease network diagram”. Nodes represented drug molecules and co- acting targets, while edges represented the interactions between nodes.
PPI Network analysis and key target identification: Based on STRING database (https://string-db.org/), we constructed a protein interaction network (Homosepiens species selected, with a confidence threshold of Highestconfli den of 0.9), and hided isolated nodes after introducing common targets. We imported the tsv format results into Cytoscape 3.10.3 software, using CytoNCA plugin to calculate the degree and betweenness of each node, and select targets with topological parameters which were more than median as core targets.
Annotations on biological functions and pathways: We utilized DAVID database (https://pubchem.ncbi.nlm.nih.gov/), performed GO functional annotation and KEGG signaling pathway enrichment analysis of co acting targets. GO functional annotations included Biological Processes (BP), Molecular Functions (MF), and Cellular Composition (CC). The results were visualized and presented on the WeChat platform.
Molecular docking verification: Through the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/), we download and obtained the spatial structure of Tolvaptan (Compound CID: 216237), from protein database(https://www.rcsb. org/), downloaded the key target protein structure. We used Pymol 3.1.5.1 software to remove water molecules and excess small molecule ligands from the target protein, performed hydrogenation, charge calculation, and other operations on the processed target protein using AutoDockTool4.2.6 software, then imported the spatial structure into Chembio3D14.0 software for energy minimization. Simultaneously we imported the receptor and ligand files into AutoDockTool software for molecular docking and calculated the binding energy, and used Pymol software to visualize the docking results of the top 4 groups with the lowest binding energy.
Results
Basic information of tolvaptan and screening of common targets of tolvaptan and ADPKD
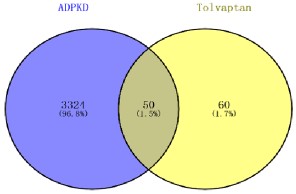
After screening, a total of 110 targets of tocilizumab and 3374 ADPKD related genes were obtained through Venn diagram analysis, 50 common target genes were identified, accounting for 1.5% of the total number of genes (Figures 1,2).
Analysis of the common target gene network of tovaptan ADPKD
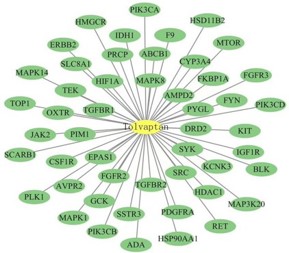
Establish a “drug target disease” interaction network using atorvastatin and 50 atorvastatin ADPKD co-acting targets. The core node is atorvastatin, and the surrounding 50 secondary nodes are potential targets for atorvastatin treatment of ADPKD. The 50 edges represent the interaction relationship between drugs and targets.
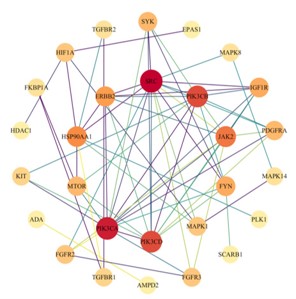
Identification of key targets in PPI network
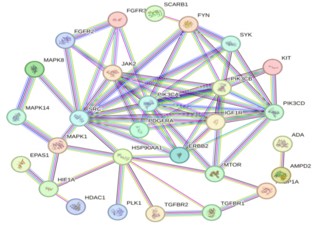
Construct a PPI network of 50 co acting targets based on the STRING database (Number of nodes: 50; Edge count: 265). After importing Cytoscape 3.9.1 software, the topology parameters were calculated using the CytoNCA plugin: The median Degree and Betweenness of each node were 4.00 and 3.00, respectively. After hiding isolated nodes, 28 key targets were obtained, and after screening, 10 core targets located in the center were obtained. The color depth of nodes represents the strength of disease correlation (increasing red gradient), and the core targets include oncogene tyrosine protein kinase (SRC), Phosphatidylinositol 4,5-diphosphate 3-Kinase catalytic subunit alpha, delta subtypes (PIK3CA and PIK3CD), Mitogen Activated Protein Kinase 14 (MAPK14), Phosphatidylinositol 4,5-diphosphate 3-Kinase Catalytic subunit beta subtype (PIK3CB), Rreceptor tyrosine protein kinase (ERBB2), etc (Figures 4,5 & Table 1).
Table 1: 10 Core targets and topological parameters of PPI network.
| Target |
Degree value |
Betweenness |
| SRC |
14 |
234.8 |
| HSP90AA1 |
8 |
209.5667 |
| MAPK1 |
4 |
67.3333 |
| PIK3CA |
13 |
55.5 |
| MTOR |
5 |
53.8 |
| FYN |
6 |
48 |
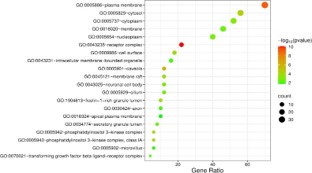
GO function enrichment analysis
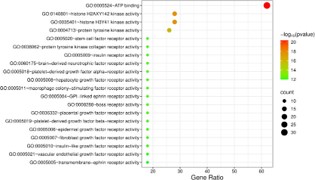
In order to analyze the key biological processes involved in the treatment of ADPKD with atorvastatin, GO functional annotation was performed on 50 common targets using the DAVID database, resulting in a total of 273 Biological Processes (BP), 37 Cellular Components (CC), 76 Molecular Functional entries (MF). BP mainly involves epidermal growth factor receptor, cell surface receptor protein tyrosine kinase, insulin-like growth factor receptor signaling pathway, etc. CC mainly involves plasma membrane, receptor complex, cytosol, etc. MF mainly involves ATP binding, histone H2AXY142 kinase activity, tyrosine kinase activity, etc. We sorted and selected the top 20 significantly enriched items based on their P-values, and generated a bubble chart using the WeChat platform. The size of the dots in the chart represents the number of targets, and the colors correspond to the P-value size (Figures 6-8).
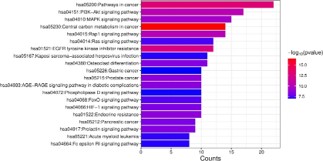
Enrichment analysis of KEGG signaling pathway
In order to explore the potential signaling pathway mechanism of tolvaptan in the treatment of ADPKD, KEGG pathway enrichment analysis was performed on 50 common target genes based on the DAVID database, and a total of 129 pathways were obtained. We selected the top 20 significant pathways and generated a bar chart using the WeChat platform, the vertical axis in the figure indicates pathway names, while the horizontal axis represents the number of enriched genes, the color gradient from blue to red reflects the magnitude of the P-value, while red corresponds to a smaller P-value. The results showed that, pathways in cancer, such as PI3K-Akt signaling pathway, MAPK signaling pathway and central carbon metabolism in cancer, the color of the item is highly enriched and redder, it indicates that PI3K-Akt, MAPK, cancer center carbon metabolism, cancer pathways and other signaling pathways are highly correlated with the pathological process of ADPKD (Figure 9).
Molecular docking verification
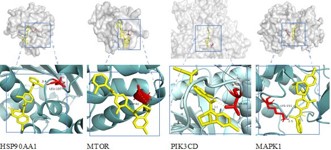
To verify the binding ability of tolvaptan to the screened core targets, we performed molecular docking with HSP90AA1, MTOR, PIK3CD, and MAPK1 proteins. The figure shows two rows of structures: the upper row represents the overall conformation of the protein, with the binding region marked with a box; The bottom row is an enlarged view of the binding site, showing the interaction between tolvaptan (yellow stick model) and target protein specific amino acids (highlighted in red), with hydrogen bonding and distance indicated. The results of this molecular docking study showed that the binding energies were all<-7.0 kcal/mol, indicating that tolvaptan forms stable binding with key targets. The lowest binding energy of -9.788 kcal/mol suggests its strong binding ability, which is likely to regulate the pathogenesis of ADPKD through interactions with these key proteins (Figure 10 & Table 2).
Table 2: Molecular docking result information table.
| Receptor |
PDBID |
Ligand |
Combining energy values |
| HSP90AA1 |
1BYQ |
Tolvaptan |
-8.218 |
| MTOR |
2GAQ |
Tolvaptan |
-7.326 |
| PIK3CD |
8S3R |
Tolvaptan |
-9.479 |
| MAPK1 |
6D5Y |
Tolvaptan |
-9.788 |
Discussion & conclusion
ADPKD is the most common hereditary kidney disease, mainly caused by mutations in the PKD1 or PKD2 genes. This disease causes structural damage and progressive decline in kidney function, and is one of the main genetic causes of End-Stage Renal Disease (ESRD). It is also accompanied by complications such as hypertension, pain, cyst bleeding, or infection. At present, the management of ADPKD mainly focuses on symptomatic treatment such as controlling blood pressure and managing complications, lacking specific therapies that can effectively delay disease progression [10]. Tovaptan, as a selective vasopressin V2 receptor antagonist, reduces the reabsorption of water by renal collecting ducts and lowers the production of Cyclic Adenosine Monophosphate (cAMP) by blocking the action of Antidiuretic Hormone (ADH). In ADPKD, reducing intracellular cAMP levels is considered a core therapeutic target because abnormally elevated cAMP promotes proliferation of cyst epithelial cells and secretion of cyst fluid [11]. This study identified 10 core targets for regulating ADPKD, including oncogene tyrosine protein kinase (SRC), phosphatidylinositol 3-kinase catalytic subunit α/β/δ (PIK3CA, PIK3CB, PIK3CD), mitogen activated protein kinase 14 (MAPK14), receptor tyrosine protein kinase (ERBB2), as well as MAPK1, MTOR, HSP90AA1, etc. SRC, as a core member of the non-receptor tyrosine kinase family, plays a pivotal role in signal transduction for cell proliferation, migration, and survival [12]. In ADPKD, SRC has been shown to be abnormally activated, which can promote excessive proliferation of cyst epithelial cells, extracellular matrix remodeling, and inflammatory response through downstream effector molecules such as phosphorylation Signal Transduction and Transcription Activator 3(STAT3), and local Focal Adhesion Kinase (FAK) [13]. The high centrality in its network is consistent with the proliferation dominant characteristics of ADPKD The PIK3 catalytic subunit is the core switch that activates the PI3K Akt pathway. The results of this study are consistent with existing literature, indicating that this pathway is activated in cyst epithelial cells. Abnormal PI3K Akt signaling plays a role by strongly inhibiting apoptosis, promoting cell cycle progression, and inducing metabolic reprogramming, such as enhancing glucose uptake, glutamine utilization, and driving continuous cyst expansion [13]. This study also enriched the PI3K dependent “cancer center carbon metabolism” pathway, further highlighting the central role of metabolic disorders in cyst growth [15]. MAPK14 and MAPK1 are important pathway nodes that regulate cellular stress response, inflammation, proliferation, and differentiation. In ADPKD, they are activated by cyst growth factors including Epidermal Growth Factor Receptor (EGFR) ligand and Tumor Necrosis Factor-α (TNF-α), which together promote sustained proliferation of cyst epithelial cells and enhance resistance to apoptosis induced signals [16]. ERBB2 serves as an important membrane receptor, its abnormal signals can simultaneously activate downstream PI3K Akt and MAPK pathways [17]. MTOR is a key downstream effector molecule of the PI3K Akt pathway, which is crucial in regulating cell growth and autophagy [18]. HSP90AA1, as a molecular partner, participates in stabilizing various kinase proteins including PI3K, SRC, AKT, etc, maintaining their active conformation and function. Its inhibition can disrupt these key proliferative signaling networks [19]. The GO enrichment analysis results showed that BP mainly involves epidermal growth factor receptors, cell surface receptor protein tyrosine kinase pathways, etc, CC mainly involves plasma membrane, receptor complexes, cell solutes, etc, MF mainly involves ATP binding, tyrosine kinase activity, etc. KEGG pathway enrichment analysis showed that the core was enriched in the PI3K-Akt signaling pathway, MAPK signaling pathway, cancer center carbon metabolism, and cancer pathways. As a vasopressin V2 Rreceptor (V2R) antagonist, the classic core mechanism of tolvaptan is to inhibit the Adenylate Cyclase (AC)-Cyclic Adenosine Monophosphate (cAMP)-Protein Kinase A (PKA) axis, reducing cystic fluid secretion. However, this study predicted through systematic analysis that tolvaptan may have the potential to target and regulate multiple key pathological signaling pathways simultaneously, not only delaying the progression of ADPKD from a purely diuretic perspective Molecular docking confirmed that tolvaptan can undergo strong molecular binding with PIK3CD. Combined with its predicted action network, such as potential effects on upstream SRC, ERBB2 or downstream MTOR, HSP90AA1, tolvaptan may directly or indirectly weaken the overall signal output of the PI3K Akt pathway [20]. This regulation will effectively counteract the strong anti-apoptotic ability and sustained proliferative effect of cyst epithelial cells caused by this pathway. Tovaptan may interact with MAPK14, MAPK1, or their upstream regulatory factor SRC [21]. Although the ultimate biological effects of the MAPK pathway are complex [22], such as c-jun N-terminal Kinase (JNK) typically promoting apoptosis [23], and Extracellular Regulated protein Kinase (ERK) promoting proliferation [24], given that MAPK pathways such as ERK have been widely reported to have significant growth promoting effects in ADPKD [25], the potential regulatory effect of tolvaptan may be manifested by inhibiting the proliferation promoting branch ERK signal or promoting the activity of specific apoptosis promoting branch JNK. This mechanism highly corresponds to the abnormal activation of EGFR receptor signaling detected in GO rich samples.
KEGG is rich in significant “cancer pathways” and “cancer center carbon metabolism” pathways, which essentially reflect the sustained activation of oncogenic metabolic reprogramming and survival signaling networks driven by downstream transcription effector molecules such as c-Myc and HIF-1 α after upstream abnormal kinase signals such as PI3K Akt and MAPK activation in ADPKD [26,27]. KEGG is rich in significant “cancer pathways” and “cancer center carbon metabolism” pathways, which essentially reflect the sustained activation of oncogenic metabolic reprogramming and survival signaling networks driven by downstream transcription effector molecules such as c-Myc and HIF-1 α after upstream abnormal kinase signals such as PI3K Akt and MAPK activation in ADPKD [26,27]. By intervening in the core kinase target groups of SRC, PI3Ks, and MAPKs, tolvaptan may indirectly inhibit these oncogenic metabolic reprogramming processes, blocking the energy and biosynthetic supply foundations that support cyst growth [28]. In summary, tolvaptan not only reduces cyst fluid secretion through V2R antagonism and cAMP inhibition effects, but may also synergistically inhibit the activity of PI3K Akt signaling pathway and MAPK signaling pathway by acting on key target networks such as SRC, PIK3CA, PIK3CB, PIK3CD, MAPK14, ERBB2, MAPK1, MTOR, HSP90AA1, and affect downstream abnormal metabolic processes such as cancer center carbon metabolism, thereby delaying the progression of ADPKD. This study provides important target clues and references for the development of novel, efficient, and highly selective ADPKD therapeutic drugs. However, the prediction results of this study are mainly based on computational models and analysis, and further in vitro and in vivo experimental verification will be a key step in elucidating these mechanisms and evaluating their therapeutic effects.
Declarations
Ethics approval and consent to participate: There is no required ethical approval or patient consent. All the data and material are available in the manuscript.
Funding: This research is funded by 2024 Shandong Province Geriatric Society Science and Technology Key Plan Project (LKJGG2024W048).
Acknowledgment: We would like to express our gratitude to all of the people who helped during the writing of this manuscript, and to the peer reviewers for their constructive opinion and suggestions.
References
- Cornec-Le Gall E, Torres VE. Genetic complexity of autosomal dominant polycystic kidney and liver diseases. J Am Soc Nephrol. 2018; 29: 13–23.
- Chebib FT, Torres VE. Assessing risk of rapid progression in autosomal dominant polycystic kidney disease and special considerations for disease-modifying therapy. Am J Kidney Dis. 2021; 78: 282–292.
- De Rechter S, Bammens B, Schaeferer F, et al. Unmet needs and challenges for follow-up and treatment of autosomal dominant polycystic kidney disease: the paediatric perspective. Clin Kidney J. 2018; 11 Suppl 1: i14–i26.
- Torres VE, Chapman AB, Devuyst O, et al. Multicenter study of long-term safety of tolvaptan in later-stage autosomal dominant polycystic kidney disease. Clin J Am Soc Nephrol. 2020; 16: 48–58.
- Kim GH. Pathophysiology of drug-induced hyponatremia. J Clin Med. 2022; 11: 5810.
- Liu Y, Pejchinovski M, Wang X, et al. Dual mTOR/PI3K inhibition limits PI3K-dependent pathways activated upon mTOR inhibition in autosomal dominant polycystic kidney disease. Sci Rep. 2018; 8: 5584.
- Reiterová J, Tesař V. Autosomal dominant polycystic kidney disease: from pathophysiology of cystogenesis to advances in the treatment. Int J Mol Sci. 2022; 23: 3317.
- Khan S, Raghuram V, Chen L, et al. Vasopressin V2 receptor, tolvaptan, and ERK1/2 phosphorylation in the renal collecting duct. Am J Physiol Renal Physiol. 2024; 326: F57–F68.
- Rigato M, Carraro G, Cirella I, et al. Effects of tolvaptan on oxidative stress in ADPKD: a molecular biological approach. J Clin Med. 2022; 11: 402.
- Zhang Y, Li Y. Progress in diagnosis and treatment of polycystic kidney disease. J Clin Intern Med. 2025; 42: 89–92.
- Niu Q, Xu Y, Ma D, et al. Research progress of drug therapy for ADPKD. J Clin Urol. 2020; 35: 842–845.
- Wang J, Zhuang S. Src family kinases in chronic kidney disease. Am J Physiol Renal Physiol. 2017; 313: F721–F728.
- Sp N, Kang DY, Joung YH, et al. Nobiletin inhibits angiogenesis by regulating Src/FAK/STAT3-mediated signaling through PXN in ER⁺ breast cancer cells. Int J Mol Sci. 2017; 18: 935.
- Margaria JP, Campa CC, De Santis MC, et al. The PI3K/Akt/mTOR pathway in polycystic kidney disease: a complex interaction with polycystins and primary cilium. Cell Signal. 2020; 66: 109468.
- Mazat JP. One-carbon metabolism in cancer cells: a critical review based on a core model of central metabolism. Biochem Soc Trans. 2021; 49: 1–15.
- Lemos FO, Ehrlich BE. Polycystin and calcium signaling in cell death and survival. Cell Calcium. 2018; 69: 37–45.
- Parker MI, Nikonova AS, Sun D, et al. Proliferative signaling by ERBB proteins and RAF/MEK/ERK effectors in polycystic kidney disease. Cell Signal. 2020; 67: 109497.
- Grahammer F, Waner N, Huber TB. mTOR controls kidney epithelia in health and disease. Nephrol Dial Transplant. 2014; 29 Suppl 1: i9–i18.
- Liu S, Xu Y, Yao X, et al. Perillaldehyde ameliorates sepsis-associated acute kidney injury via inhibiting HSP90AA1-mediated ferroptosis and pyroptosis. Int J Biol Macromol. 2025; 304: 140954.
- Wilson PD. Therapeutic targets for polycystic kidney disease. Expert Opin Ther Targets. 2016; 20: 35–45.
- Capuano I, Buonanno P, Riccio E, et al. Therapeutic advances in ADPKD: the future awaits. J Nephrol. 2022; 35: 397–415.
- Naldi L, Filbbib B, Polvani S, et al. The vasopressin receptor antagonist tolvaptan counteracts tumor growth in a murine xenograft model of small cell lung cancer. Int J Mol Sci. 2024; 25: 8402.
- Nadel G, Maik-Rachline G, Seger R. JNK cascade-induced apoptosis: a unique role in GqPCR signaling. Int J Mol Sci. 2023; 24: 13527.
- Nagao S, Yamaguchi T, Kusaka M, et al. Renal activation of extracellular signal-regulated kinase in rats with autosomal- dominant polycystic kidney disease. Kidney Int. 2003; 63: 427–437.
- Yamaguchi T, Nagao S, Wallace DP, et al. Cyclic AMP activates B-Raf and ERK in cyst epithelial cells from autosomal-dominant polycystic kidneys. Kidney Int. 2003; 63: 1983–1994.
- Song X, Pickel L, Sung HK, et al. Reprogramming of energy metabolism in human PKD1 polycystic kidney disease: a systems biology analysis. Int J Mol Sci. 2024; 25: 7173.
- Nowak KL, Hopp K. Metabolic reprogramming in autosomal dominant polycystic kidney disease: evidence and therapeutic potential. Clin J Am Soc Nephrol. 2020; 15: 577–584.
- Xia H, Huang Z, Xu Y, et al. Reprogramming of central carbon metabolism in hepatocellular carcinoma. Biomed Pharmacother. 2022; 153: 113485.